This web page was produced as an assignment for Genetics 677, an undergraduate course at UW‐Madison.
Conclusion
The focus of this project was to obtain and analyze data about BLM mined from bio-informatics tools available online. BLM is conserved mainly in eukaryotes, including human, chimpanzee, cow, mouse, rat, chicken, zebrafish, A.thaliana, rice, and P.falciparum. In general, all homologs of the BLM share DEAD/DEAH box helicase and Helicase C domains with human BLM while higher eukaryotes such as chimpanzee, cow, mouse, zebrafish, and rat share all the domains that are also present in human BLM (BLM protein domains and functions). Upon looking at BLM gene ontology, I discovered that BLM with its helicase activity involves in regulation of DNA metabolism and telomere maintenance, all of which take place in nucleus. Using String to investigate the BLM interaction network, I found that the BLM interacts with many DNA repair proteins and 2 types of telomeric proteins (TERF1 and TERF2). However, using different protein interaction database- Osprey- I found that in addition to the DNA repair proteins and telomeric proteins, BLM also interacts with WRN, the Werner Syndrome ATP-dependent helicase. These findings spurred my interest in learning about the telomeric phenotypes in a BLM patient and the protein-protein interactions between BLM, WRN, and TERF2.
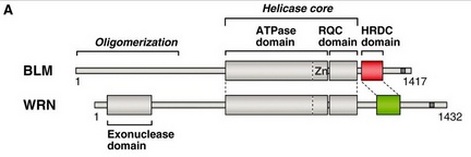
In order to find out more information about the interactions between BLM, WRN, and TERF2, I started to look into Pubmed to look for primary literature describing how all three BLM, WRN, and TERF2 play roles in a common pathway. According to Opresko et al. (2002), recent studies have shown that WRN and BLM co-localize to ALT-associated promyelocytic leukemia (PML) bodies which are localized in nucleus, but the functions of WRN and BLM in these nuclear bodies remained unclear. Alternative lengthening of telomere (ALT) is a mechanism for telomere maintenance, regulating the length and stability of telomere [5]. In short, along with many other proteins, BLM and WRN are localized to telomere by directly binding and interacting with TERF2; both BLM and WRN might play an important role in maintaining the telomere stability (To find out more about other proteins involving in ALT mechanism, please click here). Knowing that both BLM and WRN are members of RecQ helicase superfamily, I decided to look at the similarity between BLM and WRN. Although the distribution of the helicase core and HRDC domains in BLM and WRN is somehow similar, the protein sequence similarity between BLM and WRN is only 15%. Also, the N-termini of these two proteins are different; BLM protein has oligomerization near its N-terminus while WRN protein has a 3'-5' exonuclease domain. Despite the low similarity between BLM and WRN, TERF2 binds similarly to the C-terminus of either BLM or WRN [3,5].
In order to find out more information about the interactions between BLM, WRN, and TERF2, I started to look into Pubmed to look for primary literature describing how all three BLM, WRN, and TERF2 play roles in a common pathway. According to Opresko et al. (2002), recent studies have shown that WRN and BLM co-localize to ALT-associated promyelocytic leukemia (PML) bodies which are localized in nucleus, but the functions of WRN and BLM in these nuclear bodies remained unclear. Alternative lengthening of telomere (ALT) is a mechanism for telomere maintenance, regulating the length and stability of telomere [5]. In short, along with many other proteins, BLM and WRN are localized to telomere by directly binding and interacting with TERF2; both BLM and WRN might play an important role in maintaining the telomere stability (To find out more about other proteins involving in ALT mechanism, please click here). Knowing that both BLM and WRN are members of RecQ helicase superfamily, I decided to look at the similarity between BLM and WRN. Although the distribution of the helicase core and HRDC domains in BLM and WRN is somehow similar, the protein sequence similarity between BLM and WRN is only 15%. Also, the N-termini of these two proteins are different; BLM protein has oligomerization near its N-terminus while WRN protein has a 3'-5' exonuclease domain. Despite the low similarity between BLM and WRN, TERF2 binds similarly to the C-terminus of either BLM or WRN [3,5].
Therefore, these findings triggered me to ask several questions: 1) How does telomere of a Bloom's Syndrome patient look like, especially when BLM is severely truncated or non-functional? 2) In wild type cells, does BLM compete with WRN to bind to TERF2? 3) What is the expression level of WRN in a Bloom's Syndrome patient? 4) Does WRN bind more frequently to TERF2 in the absence of BLM? To address all these questions, I designed the following experiments and predicted the results of each test.
Experimental approaches:
Question 1: How does telomere of a Bloom's Syndrome patient look like, especially when BLM is severely truncated or non-functional?
Hypothesis: Extensive sister chromatid exchange can be observed while telomere sister chromatid exchange does not increase .
Reasoning: Previous studies [2] have shown that telomere in BSyn cells does not have elevated telomere sister chromatid exchange.
Question 3: What is the expression level of WRN in a Bloom's Syndrome patient?
Hypothesis: WRN is upregulated in the absence of BLM.
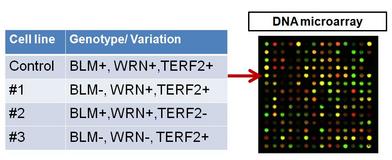
Method: Use microarray to test the expression level of WRN under different conditions. I will choose to use culture cells from Bloom's Syndrome patient as cell line #1 and use wild type human cells as control cell. Then, I will use RNAi to manipulate (knock out) certain gene in cell line#2 and cell line #3 as indicated in the genotype table below.
Hypothesis: Extensive sister chromatid exchange can be observed while telomere sister chromatid exchange does not increase .
Reasoning: Previous studies [2] have shown that telomere in BSyn cells does not have elevated telomere sister chromatid exchange.
Question 3: What is the expression level of WRN in a Bloom's Syndrome patient?
Hypothesis: WRN is upregulated in the absence of BLM.
Method: Use microarray to test the expression level of WRN under different conditions. I will choose to use culture cells from Bloom's Syndrome patient as cell line #1 and use wild type human cells as control cell. Then, I will use RNAi to manipulate (knock out) certain gene in cell line#2 and cell line #3 as indicated in the genotype table below.
Expected results: Based on the reasoning on page microarray data, I expect the level of WRN in cell line#1 will increase when BLM is absent. It is possible that WRN is highly expressed so that it can bind more frequently to TERF2 in the absence of BLM.
Question 4: Does WRN bind more frequently to TERF2 in the absence of BLM?
Hypothesis: Yes, WRN binds more frequently to TERF2.
Method: First, I will use TAP-tag and mass spectrometry to confirm the formation of TERF2-WRN complex in control cell and cell line#1. I would add GS-Tap tag to the N-terminus of WRN protein since TERF2 will be binding to the C-terminus of WRN protein. Then, I will use coimmunoprecipitation assay to find out how many of the WRN binds to TERF2. I suggest using one antibody that targets the exonuclease domain in WRN (since this is not present in BLM, we can avoid targeting TERF2-BLM complex in control cell) and another antibody that targets any part of TERF2 protein.
Expected results: After the first step, I predict that all three BLM, WRN, and TERF2 can be detected by mass spectrometry in control cell. Meanwhile, only WRN and TERF2 can be detected in cell line#1. In the coimmunoprecipitation assay, I predict that the number of TERF2-WRN complex is higher in cell line#1 compared to that of control cell.
Reasoning: Since TERF2 can bind to either BLM or WRN, so WRN needs not compete with TERF2 in the absence of BLM. Consequently, it is possible that WRN can bind more frequently to TERF2 in cells deficient of BLM than in cells with wild type BLM.
Question 4: Does WRN bind more frequently to TERF2 in the absence of BLM?
Hypothesis: Yes, WRN binds more frequently to TERF2.
Method: First, I will use TAP-tag and mass spectrometry to confirm the formation of TERF2-WRN complex in control cell and cell line#1. I would add GS-Tap tag to the N-terminus of WRN protein since TERF2 will be binding to the C-terminus of WRN protein. Then, I will use coimmunoprecipitation assay to find out how many of the WRN binds to TERF2. I suggest using one antibody that targets the exonuclease domain in WRN (since this is not present in BLM, we can avoid targeting TERF2-BLM complex in control cell) and another antibody that targets any part of TERF2 protein.
Expected results: After the first step, I predict that all three BLM, WRN, and TERF2 can be detected by mass spectrometry in control cell. Meanwhile, only WRN and TERF2 can be detected in cell line#1. In the coimmunoprecipitation assay, I predict that the number of TERF2-WRN complex is higher in cell line#1 compared to that of control cell.
Reasoning: Since TERF2 can bind to either BLM or WRN, so WRN needs not compete with TERF2 in the absence of BLM. Consequently, it is possible that WRN can bind more frequently to TERF2 in cells deficient of BLM than in cells with wild type BLM.
Future Directions
1) I would like to perform a chemical genetics experiment using a small molecule that targets and binds to the C-terminus of BLM protein. By binding the small molecule to the C-terminus of BLM, the BLM protein will be inactivated and can no longer bind to TERF2 since TERF2 binds to the C-terminus of wild type BLM. However, I realized that almost all of the small molecules that target the BLM protein binds to the BLM helicase core, but not the HRDC domain or C-terminus. Thus, I think it would be a good idea to develop a small molecule that targets the C-terminus of WRN and BLM; it could be useful in many experiments that aim to manipulate BLM or to study localization (nuclear localization signal is located near the C-terminus of BLM).
2) Since BLM and WRN have been shown to interact in vitro, it would be a good idea to investigate if TERF2-BLM complex can interact or bind directly to TERF2-WRN complex
.
3) It is clear that the role of BLM and WRN in ALT mechanism remained unclear and I think the next step should focus on determination of the functions of BLM and WRN in telomere maintenance complex.
2) Since BLM and WRN have been shown to interact in vitro, it would be a good idea to investigate if TERF2-BLM complex can interact or bind directly to TERF2-WRN complex
.
3) It is clear that the role of BLM and WRN in ALT mechanism remained unclear and I think the next step should focus on determination of the functions of BLM and WRN in telomere maintenance complex.
References:
1) S. Akiko et al. (2010). Solution structure of the HRDC domain of human Bloom syndrome protein BLM. The Journal of Biochemistry. Volume 148; Issue 4.
2) Bhattacharyya S. et al. (2010). Unwinding Protein Complexes in ALTernative Telomere Maintenance. J Cell Biochem. 109(1): 7–15. doi:10.1002/jcb.22388.
3) Lillard-Wetherell, K. et al. (2004). Association and regulation of the BLM helicase by the telomere proteins TRF1 and TRF2. Human Molecular Genetics. Vol. 13, No. 17
4) Nabetani, A., Ishikawa, F. (2010). Alternative lengthening of telomeres pathway: Recombination-mediated telomere maintenance mechanism in human cells. J Cell Biochem. 2011;149(1):5–14 doi:10.1093/jb/mvq119
5) Opresko, P. L. (2002). Telomere-binding Protein TRF2 Binds to and Stimulates the Werner and Bloom Syndrome Helicases . The Journal Of Biological Chemitsry.
2) Bhattacharyya S. et al. (2010). Unwinding Protein Complexes in ALTernative Telomere Maintenance. J Cell Biochem. 109(1): 7–15. doi:10.1002/jcb.22388.
3) Lillard-Wetherell, K. et al. (2004). Association and regulation of the BLM helicase by the telomere proteins TRF1 and TRF2. Human Molecular Genetics. Vol. 13, No. 17
4) Nabetani, A., Ishikawa, F. (2010). Alternative lengthening of telomeres pathway: Recombination-mediated telomere maintenance mechanism in human cells. J Cell Biochem. 2011;149(1):5–14 doi:10.1093/jb/mvq119
5) Opresko, P. L. (2002). Telomere-binding Protein TRF2 Binds to and Stimulates the Werner and Bloom Syndrome Helicases . The Journal Of Biological Chemitsry.