This web page was produced as an assignment for Genetics 677, an undergraduate course at UW‐Madison.
BLM Protein Domains

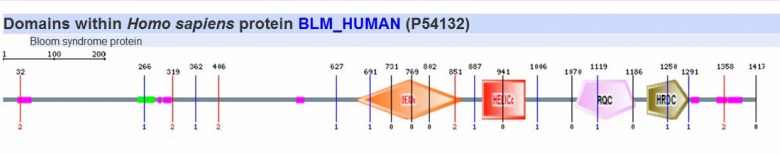
Searching on Pfam by entering the full sequence of BLM protein, I obtained a map of protein shown below. By setting Pfam-A E-value cut-off
to 1.0, Pfam found 5 Pfam-A matches to the BLM protein sequence and all the 5 matches are significant.
Functions of each domain:
The green domain represents BDHCT (NUC031) domain, a C-terminal domain in Bloom's syndrome DEAD helicase subfamily. According to Interpro database, the BDHCT domain exhibits a magnesium-dependent ATP-dependent DNA helicase activity; the helicase participates in DNA replication and repair by unwinding single- and double stranded DNA in a 3'-5' direction [1].
The red domain represents DEAD/DEAH box helicase. The DEAD box helicases unwind nucleic acids and "are involved in various aspects of RNA metabolism, including nuclear transcription, pre mRNA splicing, ribosome biogenesis, nucleocytoplasmic transport, translation, RNA decay and organellar gene expression" [1].
The blue domain is Helicase conserved C-terminal domain. This domain is found in a wide variety of helicases and helicase related proteins. Helicase conserved C-terminal domain is also described as "not an autonomously folding unit, but an integral part of the helicase" [1].
The yellow domain is RQC domain. RQC domain is a DNA-binding domain found only in RecQ family enzymes. According to Interpro database, RecQ family helicases can unwind G4 DNA, and play important roles at G-rich domains of the genome, including the telomeres, rDNA, and immunoglobulin switch regions. This domain has a helix-turn-helix structure and acts as a high affinity G4 DNA binding domain. Binding of RecQ to Holliday junctions involves both the RQC and the HRDC domains [1].
The purple domain is HRDC (Helicase and RNase D C-terminal), which has a putative role in nucleic acid binding. Mutations in the HRDC domain associated with the human BLM gene result in Bloom Syndrome (BS) [1].
The red domain represents DEAD/DEAH box helicase. The DEAD box helicases unwind nucleic acids and "are involved in various aspects of RNA metabolism, including nuclear transcription, pre mRNA splicing, ribosome biogenesis, nucleocytoplasmic transport, translation, RNA decay and organellar gene expression" [1].
The blue domain is Helicase conserved C-terminal domain. This domain is found in a wide variety of helicases and helicase related proteins. Helicase conserved C-terminal domain is also described as "not an autonomously folding unit, but an integral part of the helicase" [1].
The yellow domain is RQC domain. RQC domain is a DNA-binding domain found only in RecQ family enzymes. According to Interpro database, RecQ family helicases can unwind G4 DNA, and play important roles at G-rich domains of the genome, including the telomeres, rDNA, and immunoglobulin switch regions. This domain has a helix-turn-helix structure and acts as a high affinity G4 DNA binding domain. Binding of RecQ to Holliday junctions involves both the RQC and the HRDC domains [1].
The purple domain is HRDC (Helicase and RNase D C-terminal), which has a putative role in nucleic acid binding. Mutations in the HRDC domain associated with the human BLM gene result in Bloom Syndrome (BS) [1].
ScanProsite

The predicted intra-domain features by ScanProsite are shown below. In order to retrieve more information about post-translational modifications of the BLM protein, I entered the full BLM protein sequence and unchecked "exclude patterns with a high probability of occurrence". ScanProsite utilized different approaches such as "hits by profiles", "hits by profiles with a high probability of occurrence", and "hits by patterns" to find all motifs on BLM protein sequence.
Hits by profiles:

Figure 2. ScanProsite found three distinct profiles: Superfamilies 1 and 2 helicase ATP-binding, Helicase C-terminal, and HRDC domains, all of which were also found by Pfam. Notably, both helicase ATP-binding domain and Helicase C- terminal domain are members of helicase superfamilies 1 and 2. Unlike Pfam, ScanProsite provided more information about HRDC domain. The HRDC domain is usually found at the C-terminus of RecQ helicases. Werner Syndrome helicase (WRN), Bloom's Syndrome protein (BLM), and yeast SGS1 (the ortholog of BLM) are some of the proteins known to contain a HRDC domain [2].
Hits by profiles with a high probability of occurrence:

Figure 3. An Aspartic acid-rich region was found at amino acid 552-569 while a serine-rich region was detected near the C-terminus of BLM protein. Contained within the serine-rich region, the "Bipartite nuclear localization signal" is rich in basic amino acids such as Lysine and Arginine and it is essential to specify for the BLM protein uptake by the nucleus [2].
Hits by patterns:
By looking at consensus pattern appeared on BLM protein sequence, ScanProsite was able to identify DEAH-box subfamily ATP-dependent helicases signature domain. This domain is known to involve in ATP-dependent nucleic acid unwinding, which agreed with that described by Pfam.
Protein post-translational modifications:
It seemed like Protein Kinase C and Casein Kinase II phosphorylation sites are located throughout the full-length BLM protein. The phosphorylation by cAMP- and cGMP-dependent protein kinase occurs near N- and C-terminus while the phosphorylation by Tyrosine Kinase takes place near C-terminus of the BLM protein. Meanwhile, the BLM protein is modified by amidations and myristoylations.
Smart

Using SMART to analyze the BLM sequence, I obtained results that were slightly different from that of Pfam and ScanProsite. Even though the domain names differed from one database to another, DEAD-like helicase, helicase superfamily C-terminal, RQC and HRDC domains were also found within the BLM protein by SMART. Instead of identifying a BDHCT domain upstream of the DEAD-like helicase domain as Pfam did, SMART showed a coil coiled region and few low complexity regions. Besides listing the different domains appeared on the BLM protein, SMART provided information about type and location of mutations that will lead to Bloom's Syndrome. For example, missense mutations within the DEAD-like helicase domain and HRDC domain cause Bloom's Syndrome [3]. SMART indicated that there is no alternative splicing data for the BLM protein [3]. Interestingly, I could not find 3D structures of the BLM protein domain, but I could find those of Werner Syndrome Protein, which is also a member of RecQ helicase superfamily.
Analysis
These three databases have provided insightful information on the BLM protein. However, my favorite database website was ScanProsite because it provides additional information such as post-translational modifications and "compositional biased regions" [2]. The compositional biased regions are especially rich in a particular type of amino acid and might be important for modifications. I also liked the way SMART showed the protein map. SMART displayed the
exact intron positions in amino acid and presented them in vertical
lines, making it easy to estimate the relative position of each domain
on the BLM protein sequence.
Nonetheless, while delving into different proteomic databases to look for information about the BLM protein, I noticed that there is discrepancy in naming some domains, especially the DEAD-box helicase. But, functions of each domain remain the same in all databases. In addition, I noted that one particular domain, BDHCT, could be identified by Pfam but not by ScanProsite and SMART. The BDHCT domain might not be as significant as the other domains located downstream that ScanProsite and SMART decided not to include it in the result. For some reason, it was hard to find a 3-dimensional structure of each domain in BLM protein. In fact, the HRDC domain in the BLM protein was not solved until recent years. Click here to see the 3D structure of BLM protein domains.
Nonetheless, while delving into different proteomic databases to look for information about the BLM protein, I noticed that there is discrepancy in naming some domains, especially the DEAD-box helicase. But, functions of each domain remain the same in all databases. In addition, I noted that one particular domain, BDHCT, could be identified by Pfam but not by ScanProsite and SMART. The BDHCT domain might not be as significant as the other domains located downstream that ScanProsite and SMART decided not to include it in the result. For some reason, it was hard to find a 3-dimensional structure of each domain in BLM protein. In fact, the HRDC domain in the BLM protein was not solved until recent years. Click here to see the 3D structure of BLM protein domains.
References
1) PFAM:http://pfam.sanger.ac.uk/
2) PROSITE: http://www.expasy.ch/prosite/
3) SMART: http://smart.embl-heidelberg.de/
2) PROSITE: http://www.expasy.ch/prosite/
3) SMART: http://smart.embl-heidelberg.de/