This web page was produced as an assignment for Genetics 677, an undergraduate course at UW‐Madison.
BLM Interaction network
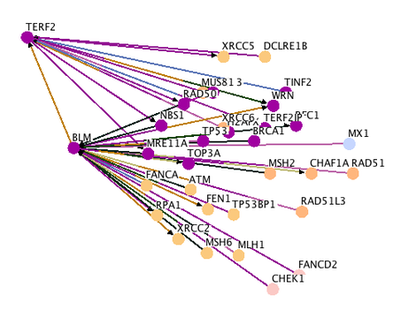
A search on String yielded an interaction network showing proteins that have been shown to interact with the BLM protein. Figure 1 shows the interactions between the BLM protein and a wide range of proteins such as telomeric binding factor (TERF1 & TERF2), DNA repair protein (RAD51), breast cancer susceptibility protein (type I and type II), and other proteins.
Analysis: I personally like String because it provides links to experiments that proved the interactions between BLM and each protein partner. This is helpful in determining the types of interaction between each protein, being either transient interaction or direct interaction. For example, for the interaction between BLM and TERF2 (as highlighted in a rectangular box in figure 1 below), the methods used by different researchers to determine BLM/TERF2 interaction included anti-bait coimmunoprecipitation assay (Association and regulation of the BLM helicase by the telomere proteins TRF1 and TRF2), enzyme linked immunosorbent assay and fluorescence microscopy assay (Telomere binding protein TRF2 binds to and stimulate the Werner and Bloom syndrome helicases), all of which indicated that BLM and TERF2 interact directly in vitro. The interaction between BLM and TERF2 struck my interest to further investigate their role in an organism, presumably in maintaining telomere stability since TERF2 is a telomeric binding factor.
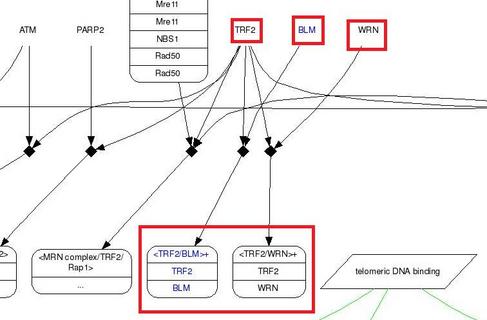
As shown in the lower panel of figure 1, I noticed that only TERF2 is linked out to Pathway Interaction Database. Pathway Interaction Database indicated that TRF2/BLM complex involves in regulation of telomerase. Also, I wanted to point out that TERF2 can bind to either BLM or WRN to form a complex (shown in Figure 2) in the telomerase regulation pathway.
Analysis: I personally like String because it provides links to experiments that proved the interactions between BLM and each protein partner. This is helpful in determining the types of interaction between each protein, being either transient interaction or direct interaction. For example, for the interaction between BLM and TERF2 (as highlighted in a rectangular box in figure 1 below), the methods used by different researchers to determine BLM/TERF2 interaction included anti-bait coimmunoprecipitation assay (Association and regulation of the BLM helicase by the telomere proteins TRF1 and TRF2), enzyme linked immunosorbent assay and fluorescence microscopy assay (Telomere binding protein TRF2 binds to and stimulate the Werner and Bloom syndrome helicases), all of which indicated that BLM and TERF2 interact directly in vitro. The interaction between BLM and TERF2 struck my interest to further investigate their role in an organism, presumably in maintaining telomere stability since TERF2 is a telomeric binding factor.
As shown in the lower panel of figure 1, I noticed that only TERF2 is linked out to Pathway Interaction Database. Pathway Interaction Database indicated that TRF2/BLM complex involves in regulation of telomerase. Also, I wanted to point out that TERF2 can bind to either BLM or WRN to form a complex (shown in Figure 2) in the telomerase regulation pathway.
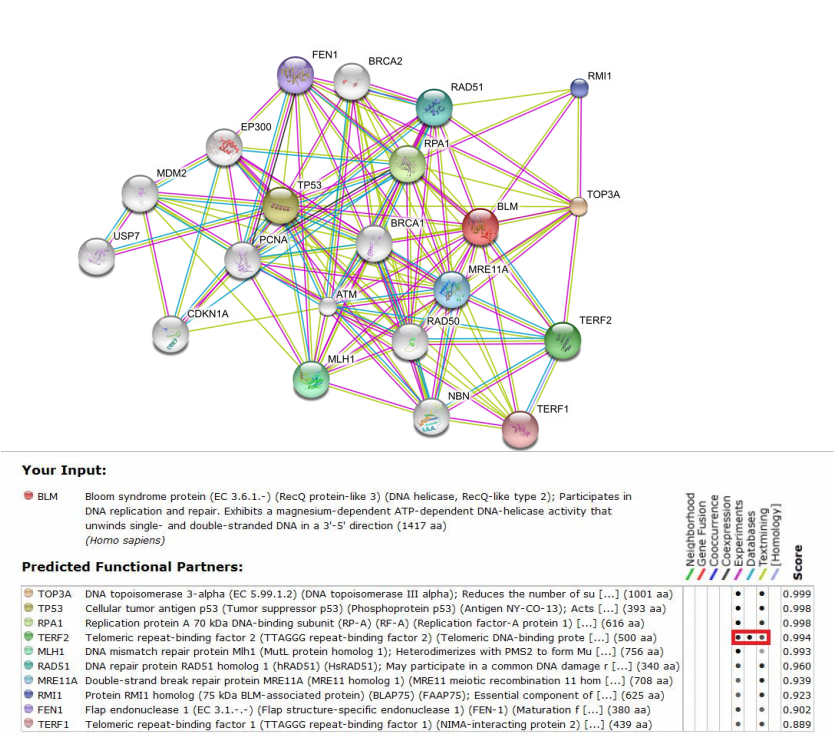
Figure 1. (Top) Interaction network of the BLM protein. (Bottom) Description of the functions of each interaction partner. STRING also provides link to experiments that showed the interactions between BLM and each protein partner. Simply click on the "circle" under the "Experiment" tab and you will be directed to corresponding experiments.
|
Figure 2. Part of the image showing TRF2/BLM and TRF2/WRN complexes in regulating telomerase. To view the entire pathway, click here.
|
|||||||
Osprey
After researching on SPRING, I decided to use Osprey to further investigate protein-protein interactions of the BLM. Osprey is an useful database to build protein interaction network as it allows users to choose from a number of model organism database and build protein network for a particular protein. I started by selecting "homo sapien database" and expanding the interaction network of BLM. Then, I choose TERF2 and expanded on its protein interaction network. Note that I did not expand protein interaction network for each BLM interaction partner because I wanted to narrow down my research to the interactions between BLM and TERF2.
Analysis: The fact that some of BLM interaction partners overlapped with those of TERF2 could be an indication that these two proteins involved in the same pathway in human. Something struck my interest was the fact that Osprey found BLM interacts with WRN, and TERF2 interacts with WRN, interactions that were not seen in the result returned from STRING. Also, interactions between these protein was found using Yeast 2-hybrid, a method that was not utilized by researchers whose work was cited by STRING. Since different databases gave different results, it was clear that interactions between BLM, TERF2, and WRN are not entirely well-characterized.
Analysis: The fact that some of BLM interaction partners overlapped with those of TERF2 could be an indication that these two proteins involved in the same pathway in human. Something struck my interest was the fact that Osprey found BLM interacts with WRN, and TERF2 interacts with WRN, interactions that were not seen in the result returned from STRING. Also, interactions between these protein was found using Yeast 2-hybrid, a method that was not utilized by researchers whose work was cited by STRING. Since different databases gave different results, it was clear that interactions between BLM, TERF2, and WRN are not entirely well-characterized.
Molecular Interaction Database (MINT)
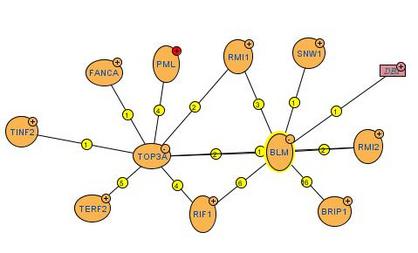
MINT presents experimentally verified protein-protein interactions obtained from the scientific literature by expert curators. By entering "BLM" under Protein name, an interaction map shown below was returned. The number on each line linking 2 proteins indicates the number of experimental evidence proving protein-protein interaction. MINT showed that there are 7 protein partners interacting with BLM protein.
Analysis: Unexpectedly, the results returned from MINT were quite different from those generated by STRING and Osprey. Except for TOP2A, the other interaction partners are unfamiliar and are generally not shown in interaction network generated by STRING and Osprey. Thus, I attempted to expand on the interaction network of TOP3A and I found that five experiments showed that TERF2 interacts with TOP3A (instead of BLM) via techniques such as anti bait coimmunoprecipitation and fluorescence microscopy. Even though recently published literature has proved that BLM directly binds to and interact with TERF2, the interaction between these 2 proteins is not represented by MINT.
Analysis: Unexpectedly, the results returned from MINT were quite different from those generated by STRING and Osprey. Except for TOP2A, the other interaction partners are unfamiliar and are generally not shown in interaction network generated by STRING and Osprey. Thus, I attempted to expand on the interaction network of TOP3A and I found that five experiments showed that TERF2 interacts with TOP3A (instead of BLM) via techniques such as anti bait coimmunoprecipitation and fluorescence microscopy. Even though recently published literature has proved that BLM directly binds to and interact with TERF2, the interaction between these 2 proteins is not represented by MINT.
BioCarta
I inputted BLM into the BioCarta, but I could not get any pathway involving BLM.
References:
1) STRING - Known and Predicted Protein-Protein Interactions
2) Breitkreutz, BJ., Stark, C., and Tyers M. (2003). Osprey: A Network Visualization System. Genome Biology,
4(3):R22.
3) Chatr-aryamontri, A., Ceol, A., Palazzi, L.M., Nardelli, G., Schneider, M.V., Castagnoli, L., and Cesareni, G.
(2007). MINT: the Molecular INTeraction database. Nucleic Acids Research, 35(Database issue):D572-D574.
2) Breitkreutz, BJ., Stark, C., and Tyers M. (2003). Osprey: A Network Visualization System. Genome Biology,
4(3):R22.
3) Chatr-aryamontri, A., Ceol, A., Palazzi, L.M., Nardelli, G., Schneider, M.V., Castagnoli, L., and Cesareni, G.
(2007). MINT: the Molecular INTeraction database. Nucleic Acids Research, 35(Database issue):D572-D574.