This web page was produced as an assignment for Genetics 677, an undergraduate course at UW‐Madison.
BLM Protein Phylogeny

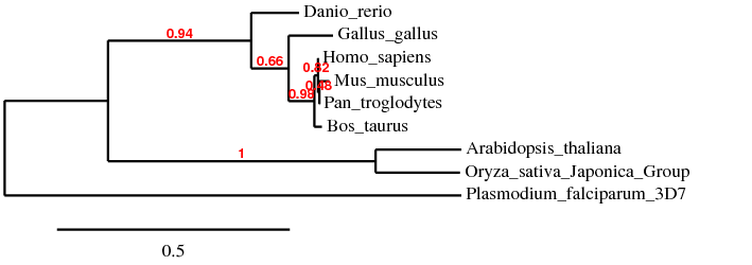
Below is a phylogenetic tree constructed via Phylogeny.fr website
Analysis
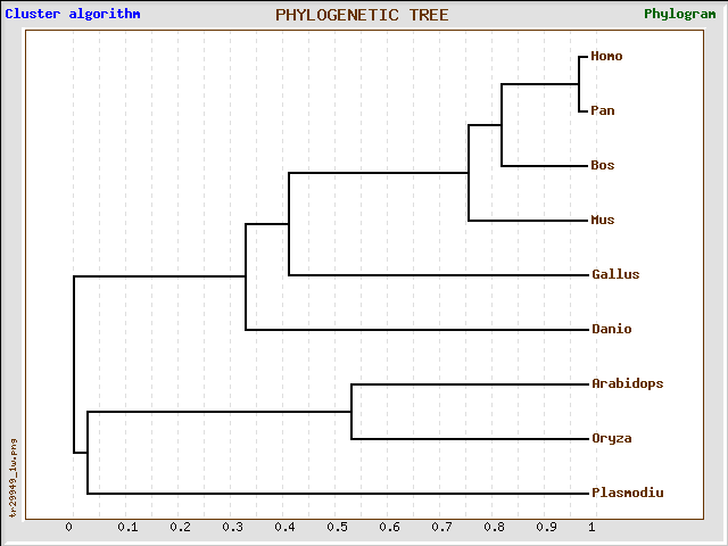
By comparing the two phylogenetic trees (shown above) from different databases, we can see that each of P. falciparum, O. sativa, A. Thaliana, D. rerio, and G. gallus has almost the same distance from branch point in both trees. On the other hand, H. sapiens, P. troglodytes, B. taurus, and M. musculus are different in their distances between branch point in both trees. In the tree generated using Phylogeny.fr website, human BLM protein is more closely related to mouse BLM protein than to that of chimpanzee. I got slightly different results even though I entered the same protein sequences into each of the program. This could be due to the different algorithm that each program uses to calculate the distance between branch points. However, as compared to the trees generated using DNA sequences, the two trees above are more useful and make more sense.
References:
1) Dereeper A., Guignon
V., Blanc G., Audic S., Buffet S., Chevenet F., Dufayard J.-F., Guindon
S., Lefort V., Lescot M., Claverie J.-M., Gascuel O. Phylogeny.fr:
robust phylogenetic analysis for the non-specialist Nucleic Acids
Research. 2008 Jul 1; 36 (Web Server Issue):W465-9. Epub 2008 Apr 19. (PubMed)
2) Phylogeny.fr: http://www.phylogeny.fr/
3) TreeTop: http://www.genebee.msu.su/services/phtree_reduced.html
2) Phylogeny.fr: http://www.phylogeny.fr/
3) TreeTop: http://www.genebee.msu.su/services/phtree_reduced.html